[wptabs] [wptabtitle]Description[/wptabtitle] [wptabcontent]
Improving Emulsion PCR libraries with Narrow Size Selections
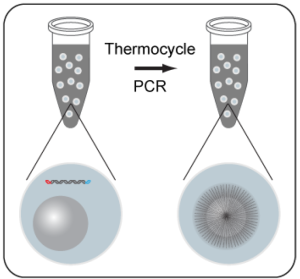
In bead-based sequencing processes, emulsion PCR is performed to generate sequencing templates on the beads. As in other PCR processes, smaller DNA fragments will be more efficiently utilized in amplification and sequencing. If the library DNA is size selected on the Pippin Prep to produce a narrow size distribution before amplification, this bias can be reduced, and libraries with more uniform sequence representation can be produced.
[/wptabcontent]
[wptabtitle]Resources[/wptabtitle] [wptabcontent]
- Brochure and Datasheets
- Presentations
- ABRF 2013: Sage - " New Automated Systems for Size-Fractionation of Protein Samples"
- AGBT 2013: Sage - "Introducing the Sage ELF: Multiple DNA size fractions from the same sample"
- AGBT 2013: Epicentre- "Gene, Transcription Start Site and Splice Site Identification Increased Through Use of Automated Preparative Gel Electrophoresis"
- AGBT 2013: PacBio (Sage Suite)- "Taking Advantage of Long Read Lengths with Improved Library Preparation Methods"
- Ion World 2012: "Faster DNA size-selection on the Pippin Prep"
- AGBT 2012: "The BluePippin System: Automated Size-selection of High Molecular Weight DNA"
- AGBT 2012: Epicentre and Sage - "Improved RNA-seq Library Quality and Workflow Enabled by Automated Preparative Gel Electrophoresis"
- AGBT 2012: Covaris and Sage - Complementary DNA Shearing and Size-selection Tools for Mate-pair Library Construction"
- PAG 2012: Digilab and Sage - "Complementary DNA Shearing and Size-selection Tools for Mate-pair Library Construction"
- AGBT 2011: "Expanded NGS Applications for the Pippin Prep DNA Size Selection System"
- AGBT 2011: Workshop Presentation, Chris Boles - "Pippin Prep 2011: New Applications - Expanded Cassette Offerings and System Improvements"
- AGBT 2010: "Automated preparative gel electrophoresis using the Pippin Prep system"
- Application Notes
- Case Studies
- Case Study: "Tag Team: Scientists Use Pippin to Optimize Nextera Libraries", Zach Herbert, Dana Farber Core Facilities
- Case Study: "Precise Sizing and SMRT Sequencing Offer Unprecedented Read Length for Clinical Studies", Robert Sebra, Mt. Sinai
- Case Study: "Better Metagenomics Methods Offer Hope for Studies with Ultra-Low DNA", Sullivan Lab, Univ. Arizona
- Case Study:" New Take on RADseq Enables High-Throughput Variant Discovery", Hoekstra Lab, Harvard Univ.
[/wptabcontent] [/wptabs]