[wptabs] [wptabtitle]Description[/wptabtitle] [wptabcontent]
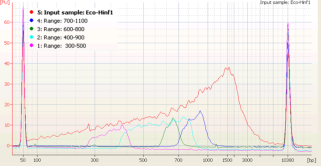
Efficient collection of broad size ranges of DNA
In range mode, the Pippin Prep can collect broad size ranges of DNA with good recovery (50-80%). This feature is useful for low input samples like those used in ChIP-seq. In addition, since collection is not dependent on detection of sample DNA, samples with DNA concentrations far below the detection limit of the Pippin Prep can be used.(click the thumbnail to enlarge).
[/wptabcontent]
[wptabtitle]Resources[/wptabtitle] [wptabcontent]
- Brochure and Datasheets
- Whitepapers
- Posters and Presentations
- ABRF 2013: Sage - " New Automated Systems for Size-Fractionation of Protein Samples"
- AGBT 2013: Sage - "Introducing the Sage ELF: Multiple DNA size fractions from the same sample"
- AGBT 2013: Epicentre- "Gene, Transcription Start Site and Splice Site Identification Increased Through Use of Automated Preparative Gel Electrophoresis"
- AGBT 2013: PacBio (Sage Suite)- "Taking Advantage of Long Read Lengths with Improved Library Preparation Methods"
- Ion World 2012: "Faster DNA size-selection on the Pippin Prep"
- AGBT 2012: "The BluePippin System: Automated Size-selection of High Molecular Weight DNA"
- AGBT 2012: Epicentre and Sage - "Improved RNA-seq Library Quality and Workflow Enabled by Automated Preparative Gel Electrophoresis"
- AGBT 2012: Covaris and Sage - Complementary DNA Shearing and Size-selection Tools for Mate-pair Library Construction"
- PAG 2012: Digilab and Sage - "Complementary DNA Shearing and Size-selection Tools for Mate-pair Library Construction"
- AGBT 2011: "Expanded NGS Applications for the Pippin Prep DNA Size Selection System"
- AGBT 2011: Workshop Presentation, Chris Boles - "Pippin Prep 2011: New Applications - Expanded Cassette Offerings and System Improvements"
- AGBT 2010: "Automated preparative gel electrophoresis using the Pippin Prep system"
- Application Notes
- Case Studies
- Case Study: "Tag Team: Scientists Use Pippin to Optimize Nextera Libraries", Zach Herbert, Dana Farber Core Facilities
- Case Study: "Precise Sizing and SMRT Sequencing Offer Unprecedented Read Length for Clinical Studies", Robert Sebra, Mt. Sinai
- Case Study: "Better Metagenomics Methods Offer Hope for Studies with Ultra-Low DNA", Sullivan Lab, Univ. Arizona
- Case Study:" New Take on RADseq Enables High-Throughput Variant Discovery", Hoekstra Lab, Harvard Univ.
[/wptabcontent] [/wptabs]